RFDiffusion – Leveraging the Power of DDPMs to Generate Protein Sequences and Structures
RFDiffusion is a new method for protein design that leverages the power of denoising diffusion probabilistic models (DDPMs) to generate protein sequences and protein structures. This approach represents a significant advance in the field of protein design, as it allows for the design of complex protein architectures and functions from simple molecular specifications.
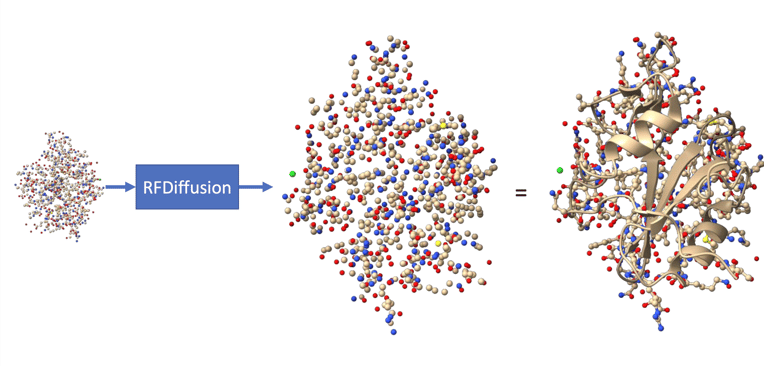
Figure 1: RFDiffusion transforms a randomly sampled point cloud of atoms into a protein structure.
Architecture and Training
RFDiffusion is based on the earlier RosettaFold (RF) architecture, similar to AlphaFold2. The authors initialize the diffusion model with the same architecture and trained weights as RosettaFold and fine-tune the model according to a diffusion modeling objective.
In contrast to RosettaFold, RFDiffusion can generate a diverse distribution of protein structures, rather than a single point estimate of protein structure.
This diffusion model has the flexibility of being able to be conditioned on any combination of sequence, secondary structure, or fold, and can generate a range of plausible structures.
After using RFDiffusion to generate a structure, the authors use the previously-released ProteinMPNN to design a sequence consistent with the designed structure.
Results
In their paper, the authors demonstrate the power and generality of their method by using it to design a range of protein architectures and functions, including monomers, binders, symmetric oligomers, enzyme active sites, and symmetric motifs. Remarkably, RFDiffusion successfully designs protein complexes with symmetries not previously observed in nature.
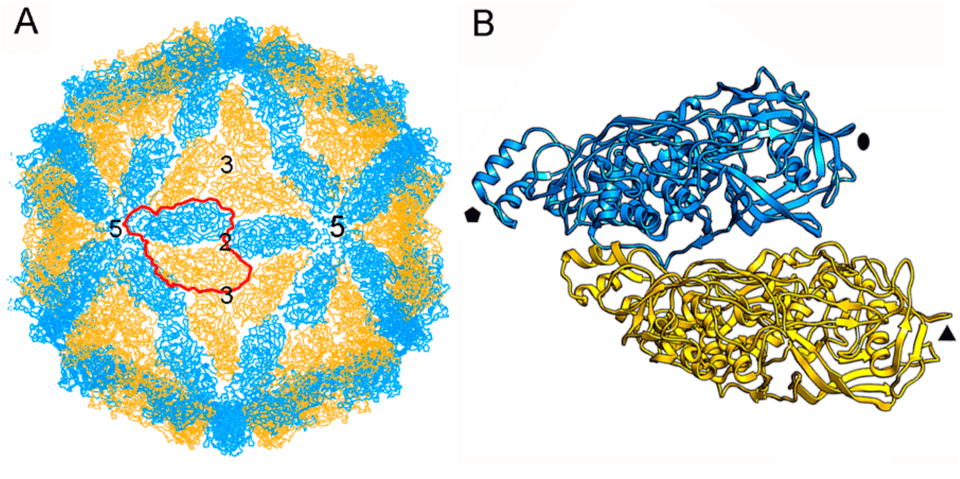
Figure 2: Example of a natural viral protein complex adopting an icosahedral symmetry. RFDiffusion was used to design proteins with previously unobserved symmetries..
The use of DDPMs for protein design represents a major advance in the field, and the RFDiffusion method is a promising approach for the design of complex protein architectures and functions.
 >
> 

