Data2Vec: A General Training Algorithm for Self-supervised Learning

What would you give to be able to train a neural network with 5% of the labels that you thought you needed? In the context of drug discovery, that could be a cost reduction of 95% for expensive experiments. Today we discuss a new paper from Meta AI, which provides a general algorithm for self-supervised learning. This algorithm bootstraps training by warm-starting the model to predict labels extracted from unlabeled data. The method is called “data2vec”.
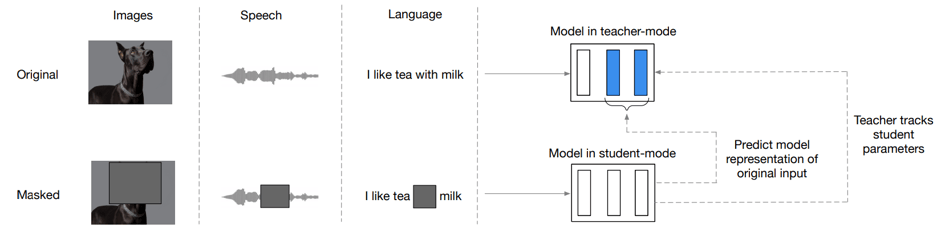
Figure 1: The data2vec training algorithm
Introduction
Representation learning is the task of finding a representation of your data that maximizes signal (which usually means information relevant to a prediction task) and minimizes noise (information which is irrelevant to a prediction task). Self-supervised learning is a type of representation learning that involves training a model using labels extracted from the unsupervised data. Data2vec is a general self-supervised training algorithm that works on all types of data.
What data modalities are common in drug discovery? Graphs are common – there are knowledge graphs used to mine the scientific literature for new drugs and targets, graphs used to model atoms and bonds in molecules, and graphs describing protein-protein and drug-protein interaction networks. 3D Euclidean data (i.e. coordinates) is also common in protein and small molecule structures. Text data in the form of scientific literature also has relevance in summarizing and querying current scientific knowledge. There is also text data in the form of medical notes. There is tabular data in CPT codes and electronic health records. There is sequence data – from the genome to the transcriptome to the proteome, not to mention string representations of small molecules (e.g. SMILES). All of these are eligible data types for data2vec.
Training
data2vec embeddings are trained “by predicting the model representations of the full input data given a partial view of the input”. The idea is that this self-supervised task will teach the model to key in on the most important features that are necessary to recreate the embedding of the masked portion. In the case of images, this might mean training to learn that a cat’s tail has the same representation as a full cat. For protein sequences, this might mean training to learn the same representation for a protein and that protein is missing its active site. This should help the neural network begin to establish the idea of concepts by forcing it to infer knowledge from partial information – without any labeled data.
The authors train a neural network with a Transformer architecture on images, speech audio, and text. There is a student model with changing weights and a target model with a moving average of the student model weights. The student model is trained to consume the masked data and produce the same representation (vector) as the representation produced by the teacher on the unmasked data. The training scheme is given in Figure 1.
The authors say that the student is trained to match the top-K blocks of the teacher. This means that we take the output of the last K layers of the teacher network for each token, average them across the K layers, and train the student network to replicate that output at its last layer for the masked tokens.
The authors refer to a time-step. This time-step refers to the position in the input sequence to the Transformer. The loss is only calculated on masked time-steps, or masked positions in the input sequence.
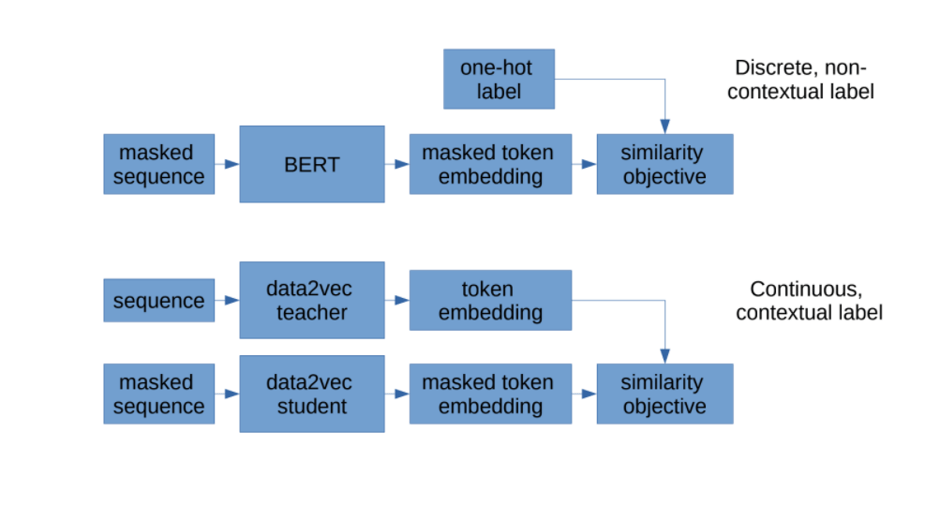
Figure 2: Comparison between BERT and data2vec, showing how data2vec learns continuous, contextual embeddings.
Continuous and Contextualized
Two key attributes of data2vec embeddings are that they are “continuous” and “contextualized”. The benefit of continuous embeddings (i.e. a real-valued vector) over discrete embeddings is that you are not restricted to a vocabulary of a fixed size – this method allows you to embed new words. Additionally, unlike original self-supervised models (e.g. BERT, see Figure 2), embeddings are contextualized. This means that embeddings can capture information about not just one token, but the tokens surrounding it. See this paragraph in a related post for more information about context in Transformers.
Results
This self-supervised pre-training task is extremely effective. To evaluate, the authors fine-tuned [with supervised learning] a model pre-trained using data2vec. Despite not being developed specifically for audio, data2vec outperforms other self-supervised methods like wav2vec. It achieves higher accuracy with one hour of audio training data than wav2vec achieves with 10h of audio training data. And that is compared to a model that already uses self-supervised learning. Compared to pure supervised learning, the model could be expected to be even more data-efficient.
This paper has tremendous implications for chemistry data. If we can leverage the enormous amount of unstructured text, chemical, and other available unlabeled data for self-supervised pre-training, we can make downstream prediction tasks much less costly in terms of labeling.
 >
> 

